¡¡Bienvenid@s a nuestra web!!
Iñaki González Restauración y Decoración es una empresa de nueva creación que surge tras el desarrollo, durante varios años, de estudios superiores en las áreas de la restauración y decoración artística.
Nuestra labor abarca distintos campos de actuación dentro de la restauración: Tratamientos de madera, limpieza por chorreo, reintegración de elementos arquitectónicos, molduras, balaustradas etc...
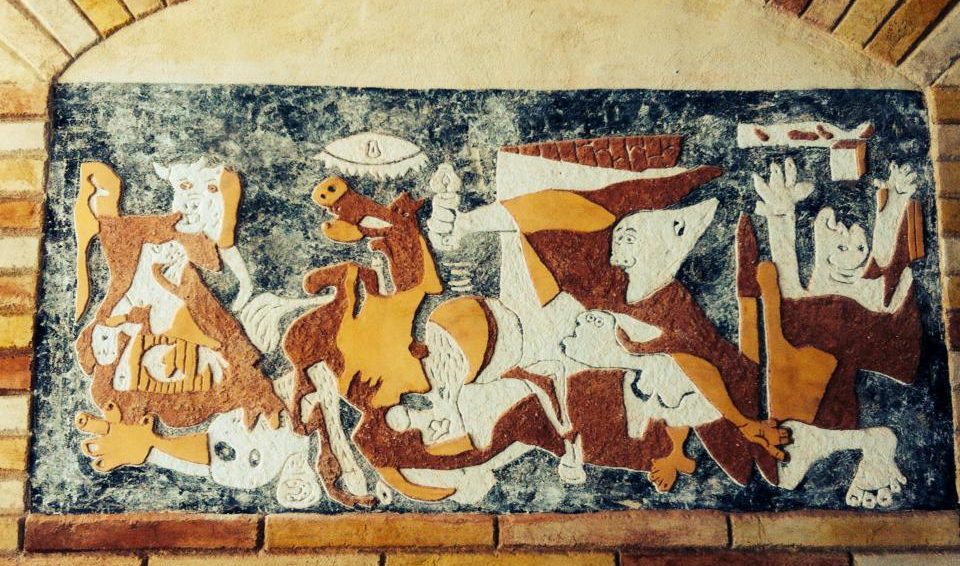
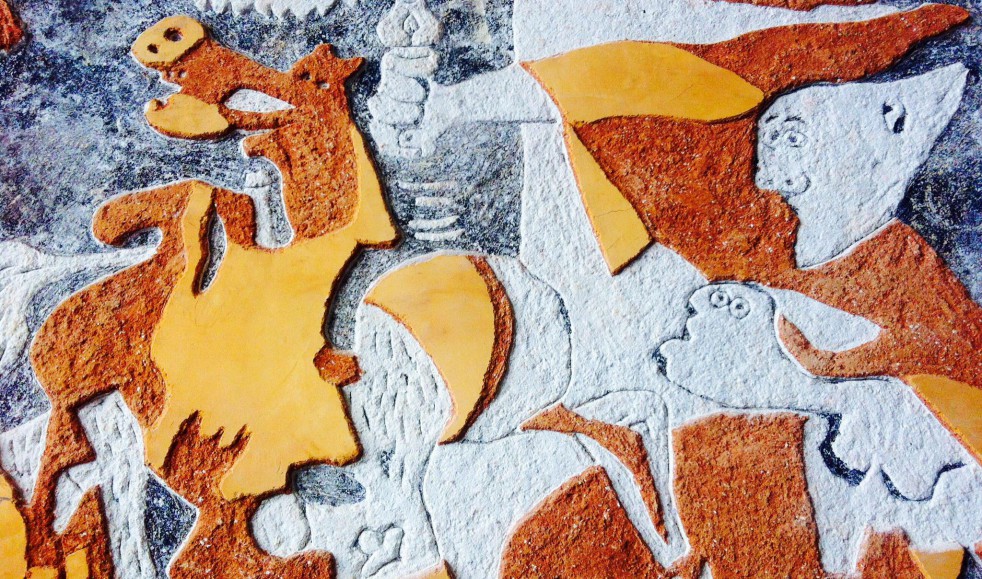
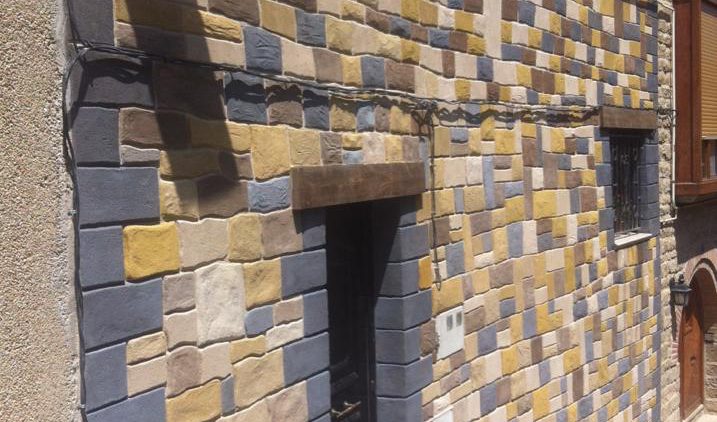
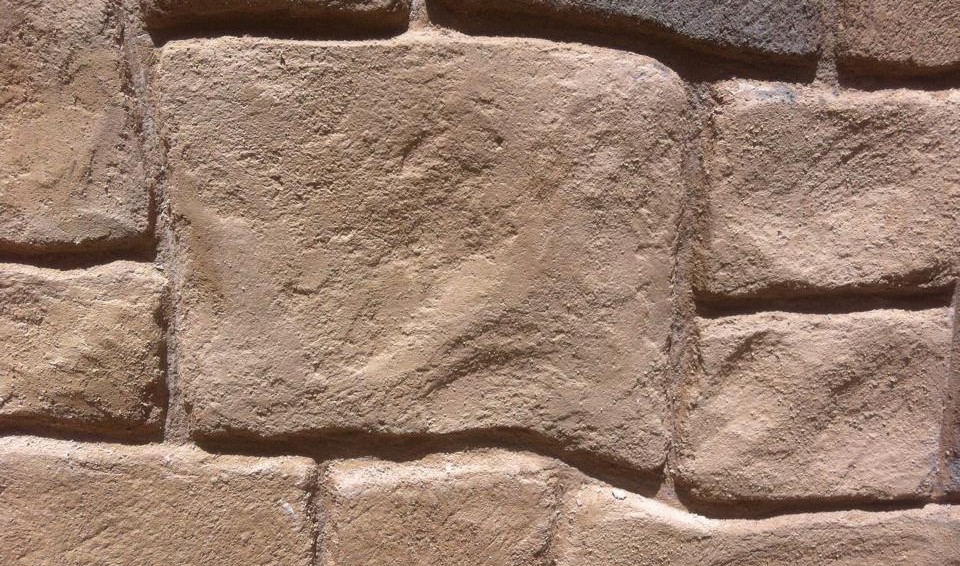
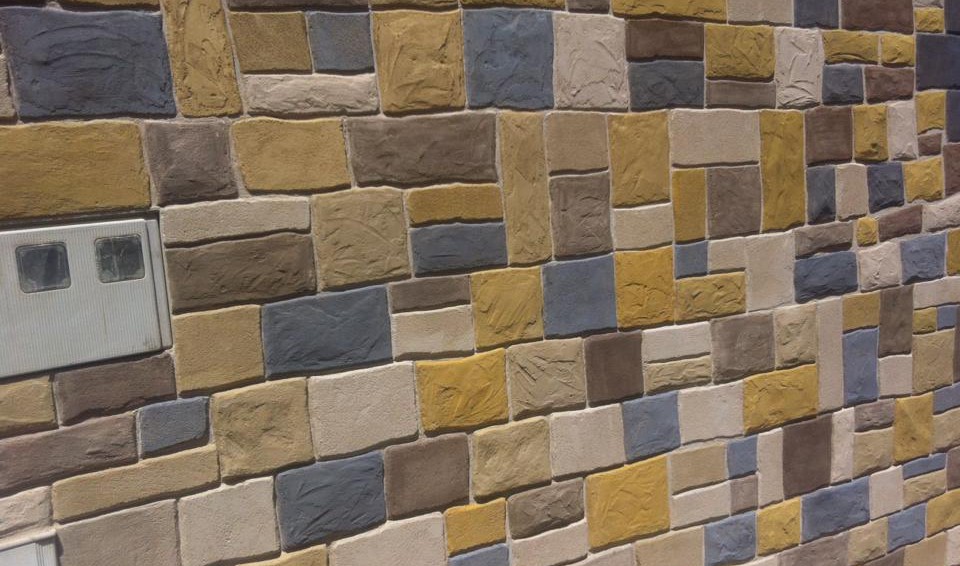
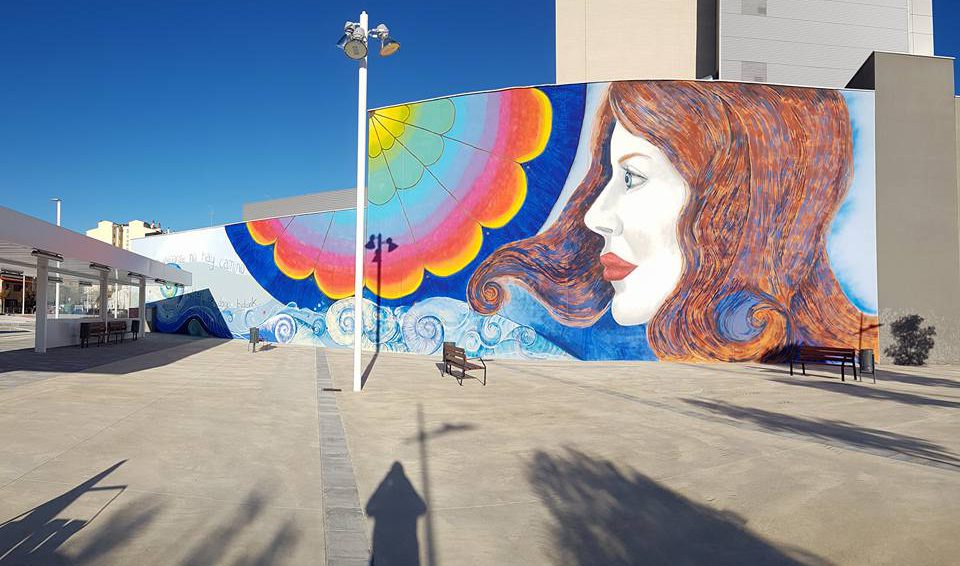
Además realizamos proyectos integrales de diseño y decoración de interiores y exteriores: Escultura, estucos, esgrafiados, imitaciones de piedra, ladrillo etc...
Consulte nuestra gama de servicios y contacte con nosotros si necesita rehabilitar o redecorar su hogar o local comercial. Presupuestos sin compromiso.